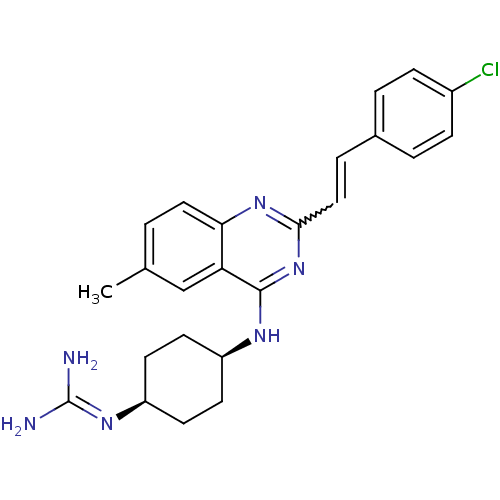
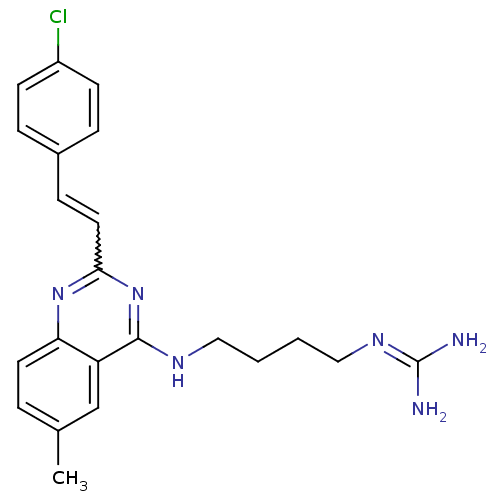
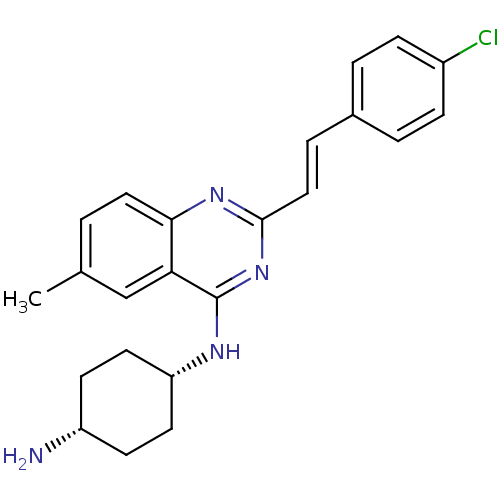
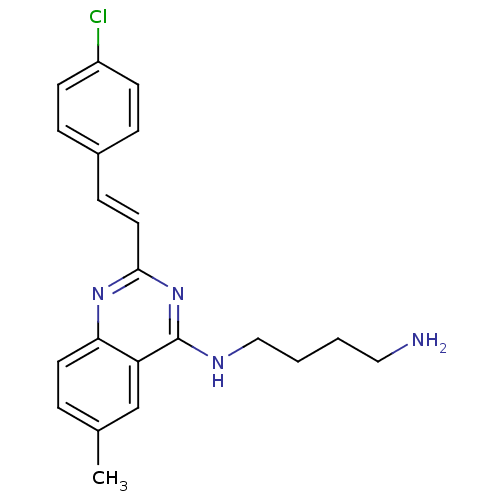
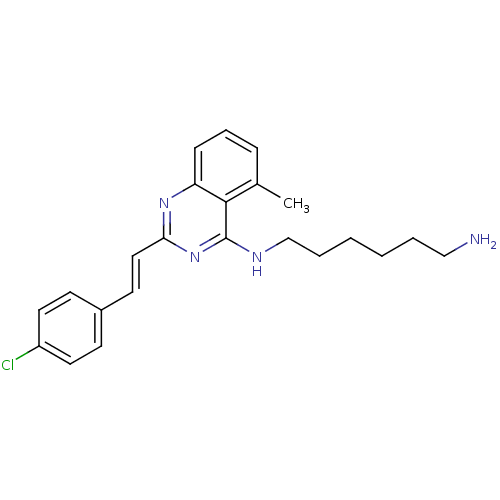
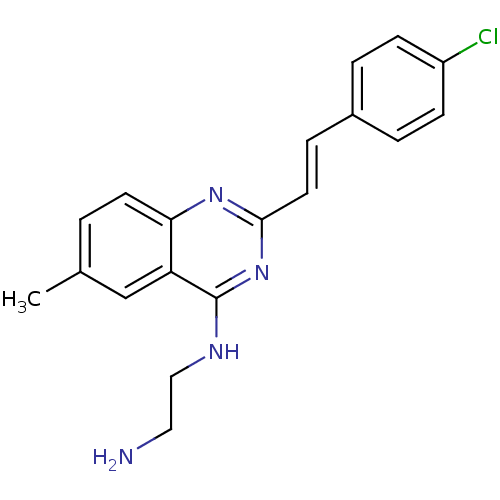
Affinity DataKi: 1.40nM ΔG°: -12.1kcal/molepH: 7.8 T: 2°CAssay Description:Competitive binding displacement analysis was performed with membranes prepared from CHO-K1 cells stably expressing receptors. After incubation, samp...More data for this Ligand-Target Pair
Affinity DataKi: 2.10nM ΔG°: -11.8kcal/molepH: 7.8 T: 2°CAssay Description:Competitive binding displacement analysis was performed with membranes prepared from CHO-K1 cells stably expressing receptors. After incubation, samp...More data for this Ligand-Target Pair
Affinity DataKi: 43nM ΔG°: -10.0kcal/molepH: 7.8 T: 2°CAssay Description:Competitive binding displacement analysis was performed with membranes prepared from CHO-K1 cells stably expressing receptors. After incubation, samp...More data for this Ligand-Target Pair
Affinity DataKi: 89nM ΔG°: -9.61kcal/molepH: 7.8 T: 2°CAssay Description:Competitive binding displacement analysis was performed with membranes prepared from CHO-K1 cells stably expressing receptors. After incubation, samp...More data for this Ligand-Target Pair
Affinity DataKi: 133nM ΔG°: -9.37kcal/molepH: 7.8 T: 2°CAssay Description:Competitive binding displacement analysis was performed with membranes prepared from CHO-K1 cells stably expressing receptors. After incubation, samp...More data for this Ligand-Target Pair
Affinity DataKi: 140nM ΔG°: -9.34kcal/molepH: 7.8 T: 2°CAssay Description:Competitive binding displacement analysis was performed with membranes prepared from CHO-K1 cells stably expressing receptors. After incubation, samp...More data for this Ligand-Target Pair
Affinity DataKi: 152nM ΔG°: -9.29kcal/molepH: 7.8 T: 2°CAssay Description:Competitive binding displacement analysis was performed with membranes prepared from CHO-K1 cells stably expressing receptors. After incubation, samp...More data for this Ligand-Target Pair
Affinity DataKi: 160nM ΔG°: -9.26kcal/molepH: 7.8 T: 2°CAssay Description:Competitive binding displacement analysis was performed with membranes prepared from CHO-K1 cells stably expressing receptors. After incubation, samp...More data for this Ligand-Target Pair
Affinity DataKi: 162nM ΔG°: -9.26kcal/molepH: 7.8 T: 2°CAssay Description:Competitive binding displacement analysis was performed with membranes prepared from CHO-K1 cells stably expressing receptors. After incubation, samp...More data for this Ligand-Target Pair
Affinity DataKi: 194nM ΔG°: -9.15kcal/molepH: 7.8 T: 2°CAssay Description:Competitive binding displacement analysis was performed with membranes prepared from CHO-K1 cells stably expressing receptors. After incubation, samp...More data for this Ligand-Target Pair
Affinity DataKi: 200nM ΔG°: -9.13kcal/molepH: 7.8 T: 2°CAssay Description:Competitive binding displacement analysis was performed with membranes prepared from CHO-K1 cells stably expressing receptors. After incubation, samp...More data for this Ligand-Target Pair
Affinity DataKi: 232nM ΔG°: -9.04kcal/molepH: 7.8 T: 2°CAssay Description:Competitive binding displacement analysis was performed with membranes prepared from CHO-K1 cells stably expressing receptors. After incubation, samp...More data for this Ligand-Target Pair
Affinity DataKi: 292nM ΔG°: -8.91kcal/molepH: 7.8 T: 2°CAssay Description:Competitive binding displacement analysis was performed with membranes prepared from CHO-K1 cells stably expressing receptors. After incubation, samp...More data for this Ligand-Target Pair
Affinity DataKi: 294nM ΔG°: -8.90kcal/molepH: 7.8 T: 2°CAssay Description:Competitive binding displacement analysis was performed with membranes prepared from CHO-K1 cells stably expressing receptors. After incubation, samp...More data for this Ligand-Target Pair
Affinity DataKi: 383nM ΔG°: -8.75kcal/molepH: 7.8 T: 2°CAssay Description:Competitive binding displacement analysis was performed with membranes prepared from CHO-K1 cells stably expressing receptors. After incubation, samp...More data for this Ligand-Target Pair
Affinity DataKi: 420nM ΔG°: -8.69kcal/molepH: 7.8 T: 2°CAssay Description:Competitive binding displacement analysis was performed with membranes prepared from CHO-K1 cells stably expressing receptors. After incubation, samp...More data for this Ligand-Target Pair
Affinity DataKi: 538nM ΔG°: -8.55kcal/molepH: 7.8 T: 2°CAssay Description:Competitive binding displacement analysis was performed with membranes prepared from CHO-K1 cells stably expressing receptors. After incubation, samp...More data for this Ligand-Target Pair
Affinity DataKi: 653nM ΔG°: -8.43kcal/molepH: 7.8 T: 2°CAssay Description:Competitive binding displacement analysis was performed with membranes prepared from CHO-K1 cells stably expressing receptors. After incubation, samp...More data for this Ligand-Target Pair
Affinity DataKi: 795nM ΔG°: -8.32kcal/molepH: 7.8 T: 2°CAssay Description:Competitive binding displacement analysis was performed with membranes prepared from CHO-K1 cells stably expressing receptors. After incubation, samp...More data for this Ligand-Target Pair
Affinity DataKi: 808nM ΔG°: -8.31kcal/molepH: 7.8 T: 2°CAssay Description:Competitive binding displacement analysis was performed with membranes prepared from CHO-K1 cells stably expressing receptors. After incubation, samp...More data for this Ligand-Target Pair
Affinity DataKi: 972nM ΔG°: -8.20kcal/molepH: 7.8 T: 2°CAssay Description:Competitive binding displacement analysis was performed with membranes prepared from CHO-K1 cells stably expressing receptors. After incubation, samp...More data for this Ligand-Target Pair